Abstract
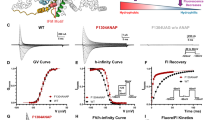
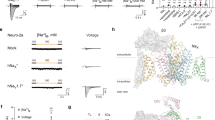
Many voltage-gated sodium channel-targeting animal peptide toxins are renowned for their potency and selectivity against insects. Understanding why these toxins selectively target insect sodium channels over their mammalian counterparts is crucial for developing safer and more effective pest control agents. Here, we present the cryoelectron microscopy (cryo-EM) structures of the insect sodium channel NavPaS bound to two naturally occurring insect-selective toxins, Av3 from the sea anemone and LqhαIT from the scorpion. Both toxins bind to the voltage-sensing domain 4 (VSD4) of NavPaS and disrupt fast inactivation by stabilizing the S4 segment in a deactivated conformation. While Av3 engages a membrane-embedded site between VSD4 and pore domain 1 (PD1), LqhαIT binds to the classical neurotoxin site 3, illustrating distinct binding modes that converge on a shared mechanism of action. These structures reveal the molecular determinants of insect selectivity and highlight the molecular coevolution of toxin-channel interactions, as corroborated by electrophysiology and toxicity assays. Leveraging these insights, we apply AI-driven protein design tools to increase the insecticidal potency of LqhαIT, resulting in a variant with a remarkable doubling in efficacy, as we confirm by insecticidal bioassays. This study illuminates the diverse mechanisms of sodium channel modulation and provides a framework for the structure-guided, AI-driven design of toxin-based biopesticides.
Similar content being viewed by others
Data availability
The cryo-EM maps have been deposited in the Electron Microscopy Data Bank (EMDB) under accession codes EMD-63108 (NavPaS-Av3); and EMD-63189 (NavPaS-LqhαIT). The atomic coordinates have been deposited in the Protein Data Bank (PDB) under accession codes 9LHZ (NavPaS-Av3); and 9LKZ (NavPaS-LqhαIT). The structures used in this paper are available in the PDB database under accession codes 1ANS, 2ASC, 5X0M, 6NT4, 7DTD, 6J8E, 7W77, 6AGF, 7DTC, 8FHD, 7WE4, 7TJ9, and 5XSY. Source data are provided with this paper.
Code availability
The ComplexDDG code has been deposited in the Zenodo database (https://doi.org/10.5281/zenodo.18150075)69. All other computational tools, including ESM-2 and A3D 2.0, are publicly available as described in references48,49.
References
Catterall, W. A. From ionic currents to molecular mechanisms: the structure and function of voltage-gated sodium channels. Neuron 26, 13–25 (2000).
Yu, F. H. & Catterall, W. A. Overview of the voltage-gated sodium channel family. Genome Biol. 4, 207 (2003).
Hodgkin, A. L. & Huxley, A. F. A quantitative description of membrane current and its application to conduction and excitation in nerve. J. Physiol. 117, 500–544 (1952).
Catterall, W. A. The molecular basis of neuronal excitability. Science 223, 653–661 (1984).
Catterall, W. A. & Swanson, T. M. Structural basis for pharmacology of voltage-gated sodium and calcium channels. Mol. Pharm. 88, 141–150 (2015).
Ahern, C. A., Payandeh, J., Bosmans, F. & Chanda, B. The hitchhiker’s guide to the voltage-gated sodium channel galaxy. J. Gen. Physiol. 147, 1–24 (2016).
Armstrong, C. M. & Bezanilla, F. Currents related to movement of the gating particles of the sodium channels. Nature 242, 459–461 (1973).
Chanda, B. & Bezanilla, F. A common pathway for charge transport through voltage-sensing domains. Neuron 57, 345–351 (2008).
West, J. W. et al. A cluster of hydrophobic amino acid residues required for fast Na(+)-channel inactivation. Proc. Natl. Acad. Sci. USA 89, 10910–10914 (1992).
Huang, J., Pan, X. & Yan, N. Structural biology and molecular pharmacology of voltage-gated ion channels. Nat. Rev. Mol. Cell Biol. 25, 904–925 (2024).
Gardill, B. R. et al. The voltage-gated sodium channel EF-hands form an interaction with the III-IV linker that is disturbed by disease-causing mutations. Sci. Rep. 8, 4483 (2018).
Wang, C. et al. Fibroblast growth factor homologous factor 13 regulates Na+ channels and conduction velocity in murine hearts. Circ. Res. 109, 775–782 (2011).
Kang, P. W. et al. Elementary mechanisms of calmodulin regulation of NaV1.5 producing divergent arrhythmogenic phenotypes. Proc. Natl. Acad. Sci. USA 118, e2025085118 (2021).
Catterall, W. A. Voltage-gated sodium channels at 60: structure, function and pathophysiology. J. Physiol. 590, 2577–2589 (2012).
Bloomquist, J. R. Ion channels as targets for insecticides. Annu. Rev. Entomol. 41, 163–190 (1996).
Soderlund, D. M. Pyrethroids, knockdown resistance and sodium channels. Pest Manag. Sci. 64, 610–616 (2008).
Silver, K. S., Song, W., Nomura, Y., Salgado, V. L. & Dong, K. Mechanism of action of sodium channel blocker insecticides (SCBIs) on insect sodium channels. Pestic. Biochem. Physiol. 97, 87–92 (2010).
Bloomquist, J. R., Coquerel, Q. R. R., Hulbert, D. & Norris, E. R. Neurophysiological action of centrally-acting spider toxin polypeptides derived from Hadronyche versuta and Tegenaria agrestis venoms. Pestic. Biochem. Physiol. 192, 105416 (2023).
King, G. F., Escoubas, P. & Nicholson, G. M. Peptide toxins that selectively target insect Na(V) and Ca(V). Channels 2, 100–116 (2008).
Bosmans, F. & Tytgat, J. Voltage-gated sodium channel modulation by scorpion alpha-toxins. Toxicon 49, 142–158 (2007).
Dong, K. et al. Molecular biology of insect sodium channels and pyrethroid resistance. Insect Biochem. Mol. Biol. 50, 1–17 (2014).
Manoleras, N. & Norton, R. S. Three-dimensional structure in solution of neurotoxin III from the sea anemone Anemonia sulcata. Biochemistry 33, 11051–11061 (1994).
Eitan, M. et al. A scorpion venom neurotoxin paralytic to insects that affects sodium current inactivation: purification, primary structure, and mode of action. Biochemistry 29, 5941–5947 (1990).
Gordon, D. et al. The differential preference of scorpion alpha-toxins for insect or mammalian sodium channels: implications for improved insect control. Toxicon 49, 452–472 (2007).
Moran, Y. et al. Molecular analysis of the sea anemone toxin Av3 reveals selectivity to insects and demonstrates the heterogeneity of receptor site-3 on voltage-gated Na+ channels. Biochem. J. 406, 41–48 (2007).
Weinberger, H. et al. Positions under positive selection-key for selectivity and potency of scorpion alpha-toxins. Mol. Biol. Evol. 27, 1025–1034 (2010).
Tugarinov, V., Kustanovich, I., Zilberberg, N., Gurevitz, M. & Anglister, J. Solution structures of a highly insecticidal recombinant scorpion alpha-toxin and a mutant with increased activity. Biochemistry 36, 2414–2424 (1997).
Rodriguez de la Vega, R. C. & Possani, L. D. Overview of scorpion toxins specific for Na+ channels and related peptides: biodiversity, structure-function relationships and evolution. Toxicon 46, 831–844 (2005).
Shen, H. et al. Structural basis for the modulation of voltage-gated sodium channels by animal toxins. Science 362, eaau2596 (2018).
Clairfeuille, T. et al. Structural basis of alpha-scorpion toxin action on Na(v) channels. Science 363, eaav8573 (2019).
Shen, H., Liu, D., Wu, K., Lei, J. & Yan, N. Structures of human Na(v)1.7 channel in complex with auxiliary subunits and animal toxins. Science 363, 1303–1308 (2019).
Xu, H. et al. Structural basis of Nav1.7 inhibition by a gating-modifier spider toxin. Cell 176, 702–715 e714 (2019).
Jiang, D. et al. Structural basis for voltage-sensor trapping of the cardiac sodium channel by a deathstalker scorpion toxin. Nat. Commun. 12, 128 (2021).
Wisedchaisri, G. et al. Structural basis for high-affinity trapping of the Na(V)1.7 channel in its resting state by tarantula toxin. Mol. Cell 81, 38–48 e34 (2021).
Neumann, B., McCarthy, S. & Gonen, S. Structural basis of inhibition of human Na(V)1.8 by the tarantula venom peptide Protoxin-I. Nat. Commun. 16, 1459 (2025).
Bende, N. S. et al. A distinct sodium channel voltage-sensor locus determines insect selectivity of the spider toxin Dc1a. Nat. Commun. 5, 4350 (2014).
Song, W., Liu, Z., Tan, J., Nomura, Y. & Dong, K. RNA editing generates tissue-specific sodium channels with distinct gating properties. J. Biol. Chem. 279, 32554–32561 (2004).
Gur Barzilai, M. et al. The specificity of Av3 sea anemone toxin for arthropods is determined at linker DI/SS2-S6 in the pore module of target sodium channels. Biochem. J. 463, 271–277 (2014).
Niklas, B. et al. Interactions of sea anemone toxins with insect sodium channel-insights from electrophysiology and molecular docking studies. Molecules 26, 1302 (2021).
Zhu, L., Gao, B., Yuan, S. & Zhu, S. Scorpion toxins: positive selection at a distal site modulates functional evolution at a bioactive site. Mol. Biol. Evol. 36, 365–375 (2019).
Li, X. et al. Structural basis for modulation of human Na(V)1.3 by clinical drug and selective antagonist. Nat. Commun. 13, 1286 (2022).
Kschonsak, M. et al. Cryo-EM reveals an unprecedented binding site for Na(V)1.7 inhibitors enabling rational design of potent hybrid inhibitors. Elife 12, e84151 (2023).
Rogers, J. C., Qu, Y., Tanada, T. N., Scheuer, T. & Catterall, W. A. Molecular determinants of high affinity binding of alpha-scorpion toxin and sea anemone toxin in the S3-S4 extracellular loop in domain IV of the Na+ channel alpha subunit. J. Biol. Chem. 271, 15950–15962 (1996).
Gordon, D. et al. Scorpion toxins affecting sodium current inactivation bind to distinct homologous receptor sites on rat brain and insect sodium channels. J. Biol. Chem. 271, 8034–8045 (1996).
Wu, F., Wu, L., Radev, D., Xu, J. & Li, S. Z. Integration of pre-trained protein language models into geometric deep learning networks. Commun. Biol. 6, 876 (2023).
Meier, J. et al. Language models enable zero-shot prediction of the effects of mutations on protein function. In Advances in Neural Information Processing Systems 34 (NeurIPS 2021).
Delgado, J., Radusky, L. G., Cianferoni, D. & Serrano, L. FoldX 5.0: working with RNA, small molecules and a new graphical interface. Bioinformatics 35, 4168–4169 (2019).
Lin, Z. et al. Evolutionary-scale prediction of atomic-level protein structure with a language model. Science 379, 1123–1130 (2023).
Pujols, J. et al. A3D 2.0 update for the prediction and optimization of protein solubility. Methods Mol. Biol. 2406, 65–84 (2022).
Li, Z., Wu, Q. & Yan, N. A structural atlas of druggable sites on Na(v). Channels Channels 18, 2287832 (2024).
Bohlen, C. J. et al. A bivalent tarantula toxin activates the capsaicin receptor, TRPV1, by targeting the outer pore domain. Cell 141, 834–845 (2010).
Yan, S., Ren, B. Y. & Shen, J. Nanoparticle-mediated double-stranded RNA delivery system: a promising approach for sustainable pest management. Insect Sci. 28, 21–34 (2021).
Nakasu, E. Y., Edwards, M. G., Fitches, E., Gatehouse, J. A. & Gatehouse, A. M. Transgenic plants expressing omega-ACTX-Hv1a and snowdrop lectin (GNA) fusion protein show enhanced resistance to aphids. Front. Plant Sci. 5, 673 (2014).
Anderson, J. A. et al. Genetically engineered crops: importance of diversified integrated pest management for agricultural sustainability. Front. Bioeng. Biotechnol. 7, 24 (2019).
Deng, S. Q., Chen, J. T., Li, W. W., Chen, M. & Peng, H. J. Application of the scorpion neurotoxin AaIT against insect pests. Int. J. Mol. Sci. 20, 3467 (2019).
Phulera, S. et al. Scorpion alpha-toxin LqhalphaIT specifically interacts with a glycan at the pore domain of voltage-gated sodium channels. Structure 32, 1611–1620 e1614 (2024).
Tan, J., Liu, Z., Nomura, Y., Goldin, A. L. & Dong, K. Alternative splicing of an insect sodium channel gene generates pharmacologically distinct sodium channels. J. Neurosci. 22, 5300–5309 (2002).
Feng, G., Deak, P., Chopra, M. & Hall, L. M. Cloning and functional analysis of TipE, a novel membrane protein that enhances Drosophila para sodium channel function. Cell 82, 1001–1011 (1995).
Zhu, Q. et al. Charge substitutions at the voltage-sensing module of domain III enhance actions of site-3 and site-4 toxins on an insect sodium channel. Insect. Biochem. Mol. Biol. 137, 103625 (2021).
Gao, R. et al. Sequence variations at I260 and A1731 contribute to persistent currents in Drosophila sodium channels. Neuroscience 268, 297–308 (2014).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Bepler, T. et al. Positive-unlabeled convolutional neural networks for particle picking in cryo-electron micrographs. Nat. Methods 16, 1153–1160 (2019).
Punjani, A., Zhang, H. & Fleet, D. J. Non-uniform refinement: adaptive regularization improves single-particle cryo-EM reconstruction. Nat. Methods 17, 1214–1221 (2020).
Zivanov, J., Nakane, T. & Scheres, S. H. W. Estimation of high-order aberrations and anisotropic magnification from cryo-EM data sets in RELION-3.1. IUCrJ 7, 253–267 (2020).
Pettersen, E. F. et al. UCSF Chimera-a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D. Biol. Crystallogr. 66, 486–501 (2010).
Liebschner, D. et al. Macromolecular structure determination using X-rays, neutrons and electrons: recent developments in Phenix. Acta Crystallogr. D. Struct. Biol. 75, 861–877 (2019).
MoleculeMind. ComplexDDG. Zenodo https://doi.org/10.5281/zenodo.18150075 (2026).
Acknowledgements
This research was funded by Fundamental and Interdisciplinary Disciplines Breakthrough Plan of the Ministry of Education of China (no. JYB2025XDXM503 to Z.Y.), the National Natural Science Foundation of China (no. 32372580 to Z.Y., no. 32260666 to S.W. and no. 92156025 to J.M.), the National Key Research and Development Program of China (no. 2025YFC3409400 to Z.Y.), and the Emerging Frontiers Cultivation Program of Tianjin University Interdisciplinary Center (to Z.Y.). We thank Jie Shen from the Tianjin Institute of Industrial Biotechnology, Chinese Academy of Sciences, for assisting with the SPR analysis.
Author information
Authors and Affiliations
Contributions
Conceptualization: H.J., and Z.Y.; Methodology: H.J., R.G., C.W., K.D., F.V.P., and Z.Y.; Investigations: H.J., R.G., C.W., H.X., S.M., Y.G., L.L., X.L., Y.L., L.Y., and R.L.; Resources: Z.Y.; Data analysis: H.J., R.G., C.W., and H.X.; Writing - original draft: H.J., R.G., C.W., H.X., Y.L., and Z.Y.; Writing - review and editing: H.J., J.X., K.D., F.V.P., Z.L., S.W., and Z.Y.; Supervision: Z.L., S.W., and Z.Y.; Project administration: Z.Y.; Funding acquisition: J.M., S.W., and Z.Y.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Kelin Xia, Nieng Yan and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jiang, H., Gao, R., Xu, H. et al. Structural insights into insect-selective sodium channel toxins drive AI-enhanced biopesticide design. Nat Commun (2026). https://doi.org/10.1038/s41467-026-70190-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-026-70190-z